
Visiting Berlin in December 2013, some friends and I spent one evening at the White Trash, a rockabilly-vibe bar decorated with hanging skeletons and a couple of conspicuous portraits of Ian “Lemmy”, the late frontman of Motörhead. Incidentally, I remember trying to guess what song was playing at a certain moment when a friend opened Shazam from her smartphone, made a short recording and presto – song, album and artist.
In the last decade, audio classification and other data-intensive statistical problems alike have been cracked with relative ease. This instalment brings together biology, physics and mathematics to create a fun deep learning application. I hope the result of nearly three months distilling the idea will be instructive and entertaining. The theme, given away by the cover figure as usual, is a familiar one. If audio classification alone seemed a good fit to the blog series, the choice of applying it to bird species identification was a no-brainer, for its accessibility and to raise awareness of the biodiversity lost to human activity. This initiative supports the Committee Against Bird Slaughter and you can help. Scroll down for more information.
By the end of this tutorial you will have trained a convolutional neural network that distinguishes among 50 bird species with up to 73.6% accuracy. The code used in this tutorial is available from this GitHub repository, and the data are hosted in this Kaggle dataset page that includes a short R notebook for guidance. Let the show begin 🂡
Introduction
Before jumping right into the analysis, let us first review some basic definitions of sound, then introduce the Fourier Transform and the dataset.
What is sound?
Sound propagates through solid, liquid and gas media as a series of pressure changes. Environmental sounds are mostly dynamic and result from the interplay of different frequencies and amplitudes that together determine pitch and loudness, respectively. The figure below depicts the transmission of a simple sound with fixed frequency and amplitude.

The human ear can pick up frequencies between 20Hz and 20,000Hz. Notably, we perceive neither frequency nor amplitude linearly, meaning a doubling of frequency or amplitude does not feel twice as high or loud, respectively – you can test this yourself here. For this particular reason, logarithmic scales such as decibel for amplitude and Mel for frequency are widely used to more closely reflect our auditory perception.
Fourier Transform
During recording, the frequencies that constitute a sound are jointly sampled to create an audio signal. The most popular technique used to work out those frequencies back from the audio signal is the Fourier Transform (FT). The FT is a calculus formula that decomposes periodic signals into the underlying frequencies, as illustrated below. For more information about the FT, here is an engaging demonstration.
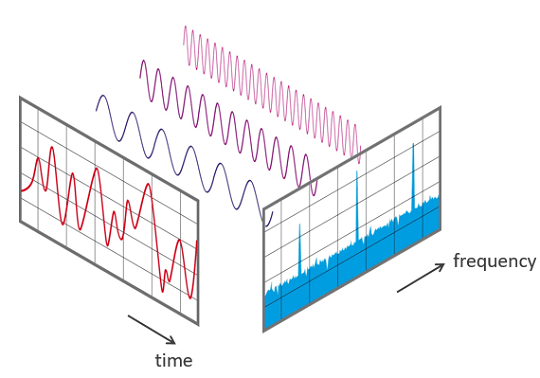
However, the classical FT only works with periodic signals, whereas most audio signals are dynamic. In the latter case, the short-time FT (STFT) can be used instead to generalise the FT over consecutive short segments of a recording, which are pre-adjusted using a window function. The resulting transformations can then be pieced together, to produce a two-dimensional spectrogram displaying intensities across the frequency spectrum and over time.
Noteworthy, the sampling rate of a recording, also measured in Hz, defines the maximum frequency that can be faithfully extracted. The elegant sampling theorem shows that any sampling rate 2N can distinguish up to a maximum frequency of N. We will see the implications of this theorem first-hand, when pre-processing the MP3 files.
Dataset
Searching for datasets of bird vocalisations in Kaggle, I found a couple of instances based on the impressive recording collection from xeno-canto, a global network of birdwatchers and enthusiasts. Unfortunately, in pursue of species classification using these datasets I could not get past 50% accuracy, partly owing to small sample sizes and recording quality, as well as confounding effects from unresolved specimen sex and vocalisation types, such as songs and calls.
A closer look into the xeno-canto portal later uncovered a convenient API that can be used to query recordings directly from the R package warbleR. This opportunity of conducting a fine-grained search with the comfort of never leaving R set the stage for designing a new dataset, using the learnings from the previous classification attempts.
The resulting dataset covers 50 bird species from 34 genera, and consists of 2150 MP3 files, totalling 8GB in size which straight away ruled out GitHub for hosting. Because you do not change a winning team, I decided to create a Kaggle dataset called Bird songs from Europe (xeno-canto) where you can find the data necessary to reproduce the reported analysis. You will also be able to query and download recordings yourself, but with no guarantee results will match. I will leave that choice up to your discretion.
To learn more about the process leading to the dataset construction, please refer to the Description section in the dataset page.
How do these recordings sound like? Take a listen to the file Cyanistes-caeruleus-405168.mp3 featuring Cyanistes caeruleus (Eurasian Blue Tit), recorded by Jarek Matusiak. You will have the chance to play more recordings later, directly from R.
Let’s get started with R
For the sake of clarity, the proposed workflow is split into two separate parts: i) download and pre-processing of bird songs, and ii) bird species classification. If your computer is underpowered, I highly recommend writing a Kaggle notebook instead.
1. Download and pre-processing of bird songs
In this first part we will use the xeno-canto API to query, filter and download a selection of bird sound recordings; then, a pre-processing routine will be set up to streamline file import, segmentation, and spectrogram extraction followed by transformation and normalisation. This undertaking will require the following packages:
parallel, to implement distributed computing during pre-processing;tidyverse, for a handful of utilities includingstringr;abind, to build input arrays;caret, for stratified train-validation-test partitioning;tuneR, to import, segment and extract spectrograms from MP3 files;warbleR, to query and download MP3 files from the xeno-canto collection.
It will also require the custom functions contained in funs.R that you may copy-paste later from the transcription, and a subdirectory mp3/ where the MP3 recordings will be stored.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Tue Feb 4 19:43:33 2020 —————————— | |
| setwd("~/Documents/Tutorials/birdsong") | |
| library(parallel) | |
| library(tidyverse) | |
| library(abind) | |
| library(caret) | |
| library(tuneR) | |
| library(warbleR) | |
| source("funs.R") | |
| # Create mp3/ if necessary | |
| if(!dir.exists("mp3/")){ | |
| dir.create("mp3/") | |
| } |
Query and download
The API query passed to warbleR::querxc() for searching the xeno-canto collection and produce the reported dataset is:
type:song type:male len_gt:30 q_gt:C area:europe
This query parameterisation returns high-quality male bird songs recorded in Europe for longer than 30 seconds. The rationale behind an Europe-wide search was to simultaneously differentiate from existing datasets and maximise species coverage; the search for male birds, which yielded ten times more results than females, to prevent sexual dimorphism from undermining results; finally, the use of a minimum recording duration to enable time-shift augmentation. These points will be reviewed in the wrap-up section.
The following code will query the xeno-canto recordings, filter entries from the query result and download the corresponding MP3 files. The filtering is based on the top 50 most frequent bird species, followed by the random downsampling of all 50 species to the least frequent. Then, the MP3 files associated with the filtered entries will be downloaded onto the newly created subdirectory mp3/. If necessary change parallel, the number of CPU cores used to speed up downloading. Besides Species, the filtered query result query includes a large amount of recording descriptors including country, long-lat coordinates, vocalisation type, quality rating and altitude, thus qualifying as supporting metadata.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| #### Download HQ male song recordings > 30s long from Europe #### | |
| query <- querxc("type:song type:male len_gt:30 q_gt:C area:europe") | |
| query$Species <- with(query, paste(Genus, Specific_epithet)) | |
| # Select top 50 most abundant bird species | |
| speciesCount <- sort(table(query$Species), decreasing = T) | |
| topSpecies <- names(speciesCount)[1:50] | |
| query <- query[query$Species %in% topSpecies, ] | |
| # Downsample to min size among the 50 classes | |
| balancedClasses <- lapply(topSpecies, function(x){ | |
| set.seed(100) | |
| sample(which(query$Species == x), min(table(query$Species))) | |
| }) %>% unlist() | |
| # Subset accordingly | |
| query <- query[balancedClasses, ] | |
| # Download using updated query | |
| querxc(X = query, download = T, path = "mp3/", parallel = 8) |
As of writing, the above procedure resulted in a total of 2150 MP3 files, 43 per species. The filenames are tagged as <GENUS>-<SPECIFIC_EPITHET>-<RECORDING_ID>.mp3, where genus and specific epithet together constitute the species name.
Pre-processing
Importing and pre-processing the MP3 files in a coordinated manner will require us to map each of the recording IDs from the metadata to the corresponding file under mp3/. To this end, we will extract the recording IDs from the filenames, match their order against Recording_ID and add their Path to the metadata query. To maximise data usability, the edited metadata table was uploaded onto the Kaggle dataset page.
The last line will play a random recording – absolutely crucial to the success of our classification task.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| #### Pre-processing #### | |
| # Read files | |
| fnames <- list.files("mp3/", full.names = T, patt = "*.mp3") | |
| # Write metadata for Kaggle dataset | |
| ids <- str_extract(fnames, pattern = "[0-9]{4,}") | |
| query$Path <- fnames[match(query$Recording_ID, ids)] | |
| write.csv(query, "metadata.csv") | |
| # Play random file – setWavPlayer in macOS if "permission denied" | |
| setWavPlayer('/usr/bin/afplay') | |
| play(sample(fnames, 1)) # esc to skip |
In order to set up the pre-processing routine, we need to carefully decide what input shape and segmentation strategy to use. In the former case, the number of spectral bands and timepoints determine the spectrogram resolution; high resolution is great, but it exerts heavy RAM and speed constraints upon training. In the latter case, if we formulate segmentation as a “sliding window” approach, the combination of parameters window size, stride and limit faces the same constraints; here we have, in addition, the trade-off between increasing sample size and reducing over-representation (e.g. long stride) or missed vocalisations (e.g. long window size).
After much trial-and-error, I picked a GPU-friendly 256 x 256 input shape and, for segmentation, a window size of 10s, a stride of 5s and a limit of 20s, therefore setting the used recording span to 30s. Below is a schematic representation of the segmentation at play, lending itself to time-shift augmentation.

As shown above, this combination of window size, stride and limit yields five samples per recording. With this plan in mind, we can now begin assembling the pre-processing routine.
The entire pre-processing will be carried out using a set of three custom functions contained in funs.R and transcribed below. In increasing order of complexity:
melspec()takes a MP3 file path, and a start and end timepoints. It first imports the MP3 file and extracts the segment between the start and end timepoints; next, it passes the resulting segment totuneR::melfcc()that will i) apply a pre-emphasis filtering, ii) compute the STFT with a Hamming window function, iii) convert the auditory frequency to the Mel scale using 256 spectral bands and 256 timepoints, and iv) take the DCT of the log-auditory-spectrum to neutralise low-energy components; the resulting 256 x 256 spectrogram is then subjected to median-based noise reduction [1] and normalisation to the maximum intensity;melslice()iteratesmelspec()over a list of MP3 files;audioProcess()employsmclapply()to leverage parallel computing ofmelslice()over a sequence of intervals defined by the user-provided window sizews,strideandlimit; next, it usesabind()to aggregate the resulting samples into a three-dimensional array; lastly, it extends the array to a single-channel dimension and reorders the four into sample x width x heigh x channel, the default channel-last Keras input format.
From the three, audioProcess() is the only that needs to be called. To better understand how pre-processing unfolds, I strongly encourage you to read the underlying code.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # read, downsample, clip, mel spec, normalize and remove noise | |
| melspec <- function(x, start, end){ | |
| mp3 <- readMP3(filename = x) %>% | |
| extractWave(xunit = "time", | |
| from = start, to = end) | |
| # return log-spectrogram with 256 Mel bands and compression | |
| sp <- melfcc(mp3, nbands = 256, usecmp = T, | |
| spec_out = T, | |
| hoptime = (end-start) / 256)$aspectrum | |
| # Median-based noise reduction | |
| noise <- apply(sp, 1, median) | |
| sp <- sweep(sp, 1, noise) | |
| sp[sp < 0] <- 0 | |
| # Normalize to max | |
| sp <- sp / max(sp) | |
| return(sp) | |
| } | |
| # iterate melspec over all samples, arrange output into array | |
| melslice <- function(x, from, to){ | |
| lapply(X = x, FUN = melspec, | |
| start = from, end = to) %>% | |
| simplify2array() | |
| } | |
| # iterate melslice over all different time windows | |
| audioProcess <- function(files, limit = 10, ws = 10, stride = 2, | |
| ncores = 8){ | |
| windowSize <- seq(0, limit, by = stride) | |
| # iterate and parallelise | |
| batches <- mclapply(windowSize, function(w){ | |
| # execute | |
| melslice(files, from = w, to = w+ws) | |
| }, mc.cores = ncores) | |
| # combine output into single array | |
| out <- abind(batches, along = 3) | |
| # reorder dimensions after adding single-channel as 4th | |
| dim(out) <- c(dim(out), 1) | |
| out <- aperm(out, c(3,1,2,4)) | |
| return(out) | |
| } |
Before kickstarting pre-processing, the data will be split into train-validation-test sets – the classifier will be trained with the train set, fine-tuned using the validation set and evaluated on the test set. To ensure meaningful representations are learned by the model, we will split our data at the file level to keep the partitions as independent as possible. For this purpose, I used a fixed stratified random split to allocate 80% of the files for training, 10% for validation and 10% for testing. This stratification is based on the species classes we need to extract from the filenames beforehand.
We can finally pass audioProcess() each of the three filename subsets along the segmentation parameters aforementioned. Reduce ncores if necessary.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Encode species from fnames regex | |
| species <- str_extract(fnames, patt = "[A-Za-z]+-[a-z]+") %>% | |
| gsub(patt = "-", rep = " ") %>% factor() | |
| # Stratified sampling: train (80%), val (10%) and test (10%) | |
| set.seed(100) | |
| idx <- createFolds(species, k = 10) | |
| valIdx <- idx$Fold01 | |
| testIdx <- idx$Fold02 | |
| # Define samples for train, val and test | |
| fnamesTrain <- fnames[-c(valIdx, testIdx)] | |
| fnamesVal <- fnames[valIdx] | |
| fnamesTest <- fnames[testIdx] | |
| # Take multiple readings per sample for training | |
| Xtrain <- audioProcess(files = fnamesTrain, ncores = 5, | |
| limit = 20, ws = 10, stride = 5) | |
| Xval <- audioProcess(files = fnamesVal, ncores = 5, | |
| limit = 20, ws = 10, stride = 5) | |
| Xtest <- audioProcess(files = fnamesTest, ncores = 5, | |
| limit = 20, ws = 10, stride = 5) |
The train-validation-test sets contain each a total of 1727, 209 and 214 MP3 files, respectively. As explained before, audioProcess() augmented those figures by five-fold, to 8635, 1045 and 1070 samples, respectively.
Once all three sets are pre-processed, we construct the corresponding responses using a one-hot encoding of the species classes. One-hot encodings can be easily computed in R using model.matrix() without an intercept.
Then, three separate do.call() instances will determine the fold-change in the train, validation and test set size due to augmentation and upsample the associated responses accordingly. Finally, all three input-response pairings are arranged into the individual lists train, val and test.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Define targets and augment data | |
| target <- model.matrix(~0+species) | |
| targetTrain <- do.call("rbind", lapply(1:(dim(Xtrain)[1]/length(fnamesTrain)), | |
| function(x) target[-c(valIdx, testIdx),])) | |
| targetVal <- do.call("rbind", lapply(1:(dim(Xval)[1]/length(fnamesVal)), | |
| function(x) target[valIdx,])) | |
| targetTest <- do.call("rbind", lapply(1:(dim(Xtest)[1]/length(fnamesTest)), | |
| function(x) target[testIdx,])) | |
| # Assemble Xs and Ys | |
| train <- list(X = Xtrain, Y = targetTrain) | |
| val <- list(X = Xval, Y = targetVal) | |
| test <- list(X = Xtest, Y = targetTest) |
We conclude this first part with the visualisation of a spectrogram randomly drawn from the train set, along with a handful reference frequencies in Mel scale. Before moving on to the second part, make sure that the three pre-processed sets train, val and test are saved into prepAudio.RData (~2.6GB in size).
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Plot spectrogram from random training sample – range is 0-22.05 kHz | |
| image(train$X[sample(dim(train$X)[1], 1),,,], | |
| xlab = "Time (s)", | |
| ylab = "Frequency (kHz)", | |
| axes = F) | |
| # Generate mel sequence from Hz points, standardize to plot | |
| freqs <- c(0, 1, 5, 15, 22.05) | |
| mels <- 2595 * log10(1 + (freqs*1e3) / 700) # https://en.wikipedia.org/wiki/Mel_scale | |
| mels <- mels – min(mels) | |
| mels <- mels / max(mels) | |
| axis(1, at = seq(0, 1, by = .2), labels = seq(0, 10, by = 2)) | |
| axis(2, at = mels, las = 2, | |
| labels = round(freqs, 2)) | |
| axis(3, labels = F); axis(4, labels = F) | |
| #### Save #### | |
| save(train, val, test, file = "prepAudio.RData") |

The default maximum frequency set by melfcc() upon extraction is, I hope familiarly, half the recording sampling rate. The attentive reader might wonder if all recordings possess the same sampling rate of 44.1kHz, as hinted from the above spectrogram. Regrettably not, meaning spectrograms might display different frequency spans, and consequently patterns shifted or stretched vertically. This issue will also be reviewed in the wrap-up section.
Now indulge yourself with a well-deserved coffee break! ☕️
2. Bird species classification
In this second part we will design, train and evaluate the convolutional neural network (CNN) classifier using Keras. Before dealing with any of that, a few things need to be sorted out. Firstly, we import the following packages from a new R session:
keras, to design, compile and fit the CNN;caretande1071, to build the confusion matrices;pheatmap, to visualise the confusion matrices;RColorBrewer, to create a colour palette.
Then, we activate a Keras backend. My backend, as indicated in the GitHub repository page, is a GPU configuration of plaidML from a Conda environment called plaidml. If you prefer using TensorFlow instead, make the necessary changes to the code. Lastly, we load prepAudio.RData to bring back our pre-processed train, val and test sets.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Fri Feb 7 15:49:46 2020 —————————— | |
| setwd("~/Documents/Tutorials/birdsong") | |
| library(keras) | |
| use_condaenv("plaidml") | |
| use_backend("plaidml") | |
| k_backend() # plaidml | |
| library(tidyverse) | |
| library(caret) | |
| library(e1071) | |
| library(pheatmap) | |
| library(RColorBrewer) | |
| # Read processed data | |
| load("prepAudio.RData") |
Model architecture
A CNN is by no means the most effective approach to audio classification. In fact, borrowing the basis of computer vision applications to classify audio data might seem odd at first. It is, however, sufficiently powerful and simple for the scope of a tutorial. If you need some background on CNNs, have a look into this past tutorial.
Assuming the different species use different frequency ranges, as suggested from the R notebook analysis, I hypothesised that an optimal classifier should learn song representations that are equivariant to time, but not frequency. This conundrum of two-dimensional convolution using audio data led Goodfellow and colleagues [2] to similar conclusions. Therefore, I suggest using a two-dimensional CNN model that learns pseudo-spatial representations and later projects the feature map over time to keep track of frequency ranges. The diagram below summarises the audio classification workflow and provides a detailed overview of the proposed model architecture.

The top part of the diagram represents the acquisition and pre-processing of the audio data, resulting in the three-dimensional input arrays we produced earlier.
The bottom part is a deconstruction of the CNN that consists of four two-dimensional convolution rounds, each followed by max-pooling, leading to a pair of fully-connected layers, the last of which returning all 50 class probabilities. This CNN architecture uses dropout between layers and 3 x 3 convolution kernels throughout. The most unusual feature, highlighted in red, is the last max-pooling that projects the feature map over time. The resulting projections are subsequently “flattened” into a layer of size . This layer is then fully connected to a layer of size 128, and this in turn with the output layer, of size 50. At last, the softmax transformation of the output from this last layer computes the class probabilities. The largest of the 50 probabilities, represented by a black output unit, determines the species predicted.
Model training
At this point we have ready-to-use data and a model architecture, the key requirements to implement, compile and train a Keras model. Writing down the desired CNN structure to R code is made straightforward by the use of magrittr pipes.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Build model | |
| model <- keras_model_sequential() %>% | |
| layer_conv_2d(input_shape = dim(train$X)[2:4], | |
| filters = 16, kernel_size = c(3, 3), | |
| activation = "relu") %>% | |
| layer_max_pooling_2d(pool_size = c(2, 2)) %>% | |
| layer_dropout(rate = .2) %>% | |
| layer_conv_2d(filters = 32, kernel_size = c(3, 3), | |
| activation = "relu") %>% | |
| layer_max_pooling_2d(pool_size = c(2, 2)) %>% | |
| layer_dropout(rate = .2) %>% | |
| layer_conv_2d(filters = 64, kernel_size = c(3, 3), | |
| activation = "relu") %>% | |
| layer_max_pooling_2d(pool_size = c(2, 2)) %>% | |
| layer_dropout(rate = .2) %>% | |
| layer_conv_2d(filters = 128, kernel_size = c(3, 3), | |
| activation = "relu") %>% | |
| layer_max_pooling_2d(pool_size = c(28, 2)) %>% | |
| layer_dropout(rate = .2) %>% | |
| layer_flatten() %>% | |
| layer_dense(units = 128, activation = "relu") %>% | |
| layer_dropout(rate = .5) %>% | |
| layer_dense(units = ncol(train$Y), activation = "softmax") |
Next, a call to summary() prints out the architecture and parameterisation – make sure everything looks right. We then proceed with compiling the model using the Adam optimiser, with a learning rate and decay of and
, respectively, and a categorical cross-entropy loss. Finally, the model is fit using a batch size of 16 over a total of 50 epochs. The validation set
val is passed as validation_data to facilitate model fine-tuning.
Once training is over, a plot(history) call will bring up an overview of loss and accuracy over the epochs in both train and validation sets. The resulting model object can then be saved as a HDF5 file for distribution or deployment.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Print summary | |
| summary(model) | |
| model %>% compile(optimizer = optimizer_adam(decay = 1e-5), | |
| loss = "categorical_crossentropy", | |
| metrics = "accuracy") | |
| history <- fit(model, x = train$X, y = train$Y, | |
| batch_size = 16, epochs = 50, | |
| validation_data = list(val$X, val$Y)) | |
| plot(history) | |
| # Save model | |
| # model %>% save_model_hdf5("model.h5") |

On my end, training ran just under one hour at 68s per epoch. The reported training and validation accuracies reached ~90% and ~75%, respectively.
Model evaluation
We conclude this tutorial with the inspection of confusion matrices to more closely investigate accuracy from the validation and test set predictions. Confusion matrices are contingency tables that count each predicted class (row-wise) given the observed class (column-wise). As a result, for a k-class problem there is a confusion matrix whose diagonal entries capture successful predictions.
To this end, we first extract the predicted probabilities predProb and use a simple apply() trick to retrieve, from every sample, the class associated with the maximum probability, predClass. Next, we pass the predicted and observed classes to confusionMatrix() and finally visualise the resulting $table object via pheatmap(). Accuracy can be easily estimated by computing the average number of matches across predClass and trueClass.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Grep species, set colors for heatmap | |
| speciesClass <- gsub(colnames(train$Y), pat = "species", rep = "") | |
| cols <- colorRampPalette(rev(brewer.pal(n = 7, name = "RdGy"))) | |
| # Validation predictions | |
| predProb <- predict(model, val$X) | |
| predClass <- speciesClass[apply(predProb, 1, which.max)] | |
| trueClass <- speciesClass[apply(val$Y, 1, which.max)] | |
| # Plot confusion matrix | |
| confMat <- confusionMatrix(data = factor(predClass, levels = speciesClass), | |
| reference = factor(trueClass, levels = speciesClass)) | |
| pheatmap(confMat$table, cluster_rows = F, cluster_cols = F, | |
| border_color = NA, show_colnames = F, | |
| labels_row = speciesClass, | |
| color = cols(max(confMat$table)+1)) | |
| # Accuracy in validation set | |
| mean(predClass == trueClass) # 0.7541 |

In line with the training history, we see that the validation set accuracy was 75.4%. We will now turn our attention to the test set and apply the same recipe.
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| # Test set prediction | |
| predXProb <- predict(model, test$X) | |
| predXClass <- speciesClass[apply(predXProb, 1, which.max)] | |
| trueXClass <- speciesClass[apply(test$Y, 1, which.max)] | |
| # Plot confusion matrix | |
| confMatTest <- confusionMatrix(data = factor(predXClass, levels = speciesClass), | |
| reference = factor(trueXClass, levels = speciesClass)) | |
| pheatmap(confMatTest$table, cluster_rows = F, cluster_cols = F, | |
| border_color = NA, show_colnames = F, | |
| labels_row = speciesClass, | |
| color = cols(max(confMatTest$table)+1)) | |
| # Accuracy in test set | |
| mean(predXClass == trueXClass) # 0.7364 | |
| # Write sessioninfo | |
| writeLines(capture.output(sessionInfo()), "sessionInfo") |
The test set accuracy was 73.6%, very close to the validation set accuracy and reflecting a low generalisation error.

Taking into account the inclusion of representatives from common genera, one could expect misclassification among close species to be greater than among distant species. The predictions, however, show no discernible pattern in this respect.
Wrap-up
We successfully trained a CNN classifier that distinguishes among 50 bird species from their vocalisations with an accuracy of 73.6%. This concluding section reviews some of the options exercised and spreads out ideas for future work:
- Input data – the data used to train the CNN classifier consisted of male bird songs recorded in Europe. Provided the right conditions and particularly sample size, this approach can be extended to female birds and other regions. On the other hand, I suspect other vocalisation types will not be as discriminative of species as songs;
- Sampling rate – the MP3 files have different sampling rates that might distort the spectrograms. This problem can be easily fixed by passing a
maxfreqtomelfcc()or usingtuneR::downsample()on the imported MP3 object. In any case, here is a breakdown of the files by sampling rate – 48kHz (1141), 44.1kHz (969), 32kHz (27), 24kHz (1), 22.05kHz (7), 16kHz (4), 8kHz (1). The lion share goes to 48kHz and 44.1kHz (~98%); - Augmentation – recording time is the bottleneck in time-shift augmentation, as querying files longer than 30s returns much fewer results. The usage of a window size of 10s as suggested elsewhere worked better compared to 5s and 8s. Setting a stride to shorter than 5s increased sample size considerably, alas at the expense of RAM exhaustion. Other types of augmentation are yet to be tested;
- Mel frequency bands – keeping as many as 256 Mel bands might be counterproductive. Using fewer bands could prove important to maintain performance and free up RAM for larger sample sizes;
- High-pass filter – following accounts of the successful reduction of environmental noise [1], a high-pass filter of 1kHz was initially added to the pre-processing routine, using the
minfreqargument frommelfcc(). However, some pre-processed spectrograms had missing bird songs as can be deduced from the R notebook analysis. Interestingly, the removal of the filter did not affect accuracy, suggesting the model picks up resonating higher frequencies from low-pitch bird songs; - Spectrogram (de)noising – in attempt to prevent the CNN from memorising overlaps between augmented spectrograms, Gaussian-noise addition was initially implemented in the pre-processing routine; this, however, did not materialise into performance. Conversely, the introduction of spectrogram median-based noise reduction [1] improved validation accuracy by ~5%;
- Image analysis – image analysis techniques, including intensity thresholding followed by morphological closing [3] were successfully applied to spectrogram pre-processing. This kind of cross-disciplinary interventions substantially expands the available toolkit;
- Model architecture – the simple model architecture used here can be easily adapted and modified. Curiously, switching from batch normalisation to dropout layers boosted training speed and performance. Moreover, the application of 5 x 5 kernels on the top convolutional layers [4] or stepwise, asymmetric max-pooling with size 4 x 2 [5] and 3 x 2 proved unfruitful;
- Transfer learning – the convolutional base of the VGG16 CNN was used with a three-channel version of the input data, turning out to be slow and largely inaccurate; since simpler models had proved effective, pre-trained networks were dismissed.
Please support
Well over a thousand birds were part of this story. What better way to pay tribute than to help protecting them? This tutorial supports the Committee Against Bird Slaughter (CABS) – an international NGO committed to protecting wild birds from illegal poaching throughout the migratory flyways of the Mediterranean. You can read more about the organisation here. If you enjoyed this content, I urge you to support their work.

Committee Against Bird Slaughter
The Committee Against Bird Slaughter (CABS) is a charitable Non-Governmental Organisation working to protect migratory birds from illegal poaching and over exploitation. We are an activist association with a lean administration based from our Head office in Bonn, Germany.
€1.00
I want to thank Lloyd Scott for introducing the CABS and the xeno-canto community for amassing and sharing such a large collection of bird sound recordings. My thoughts are with those afflicted by the COVID-19 outbreak, stay safe. Finally, if you have any questions, corrections, suggestions or general remarks please leave a comment below. See you next time 🂡

References
[1] Stowell, D and Plumbley, MD (2014). Automatic large-scale classification of bird sounds is strongly improved by unsupervised feature learning. PeerJ 2:e488; DOI 10.7717/peerj.488
[2] Goodfellow, I et al., (2017). Deep Learning. Chapter 9, p349
[3] Khal, S et al., (2017). Large-Scale Bird Sound Classification using Convolutional Neural Networks. Working notes on CLEF
[4] Sprengel, E et al., (2016). Audio Based Bird Species Identification using Deep Learning Techniques. CLEF
[5] Salamon, J and Bello, JP (2017). Deep Convolutional Neural Networks and Data Augmentation for Environmental Sound Classification. IEEE Signal Processing Letters 24, 3, 279–283
Citation
de Abreu e Lima, F (2020). poissonisfish: Audio classification in R. From https://poissonisfish.com/2020/04/05/audio-classification-in-r/

Great post! Do you know if wavelets can be use for this type of analysis?
LikeLike
Hi Victor, thanks for the kind words. I suppose yes, the STFT used here could be adjusted / replaced accordingly. Best, Francisco
LikeLike
Thanks a lot for sharing great knowledge. I am working on detecting insect pest sounds in stored grains to prevent losses, this is indeed useful.
LikeLike
Hi Carlito, glad you found it helpful! Good luck with your project, it sounds interesting
LikeLike